1. From Linked Lists to Graphs
A Graph is a collection of nodes (vertices) connected by edges.
- Linked List as Graph: A simple graph where nodes are connected in a linear sequence.
- Singly-Linked: Each node has one “forward” edge.
- Doubly-Linked: Each node has a “forward” and a “backward” edge.
- Tree as Graph: A more complex graph structure that is connected and contains no undirected cycles.
2. Defining a Tree
A graph is a tree if it meets these equivalent criteria:
- It is connected and has no undirected cycles.
- It is acyclic, but adding any single edge creates a cycle.
- It is connected, but removing any single edge makes it disconnected.
- There is a unique simple path between any two vertices.
- For vertices, the tree has exactly edges.
Note on Edge Cases: An “empty” (null) tree with 0 nodes and a tree with a single node are both considered valid trees in data structure contexts.
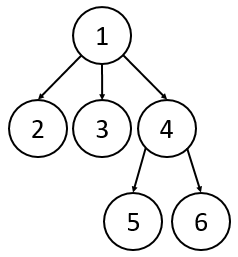
3. Rooted vs. Unrooted Trees
Rooted Trees (Hierarchical)
Rooted trees have a clear top-down structure defined by parent-child relationships.
- Root: The single top node with no parent.
- Parent/Child: Direct vertical relationships.
- Ancestors/Descendants: All nodes along the path upward or downward.
- Leaves: Nodes with no children.
- Internal Nodes: Nodes that have at least one child.
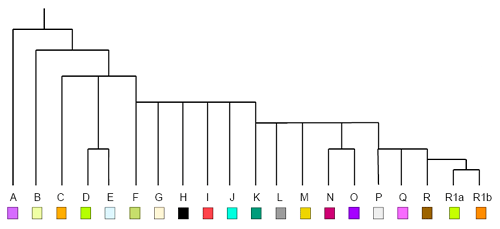
Y-Chromosome Evolutionary Trees
In Bioinformatics, researchers study paternal lineages by constructing evolutionary trees from Y chromosomes. Because the Y chromosome is passed directly from father to son as a single DNA sequence, it creates a clear hierarchical record.
- Directionality: The tree represents time in a “top-down” fashion. Traversing from the root toward the leaves move you forward in time.
- The Root: In this context, the root node represents the Most Recent Common Ancestor (MRCA) of all the individuals in the dataset.
- The Leaves: These represent the specific individuals whose DNA was sequenced.
Unrooted Trees (Relational)
Unrooted trees lack a hierarchy or a specific starting “root.”
- Neighbors: Any nodes directly connected by an edge.
- Leaf: A node with exactly one neighbor.
- Internal Node: A node with more than one neighbor.
- Application: Used in Bioinformatics to show biological similarity between species when the common ancestor is unknown.
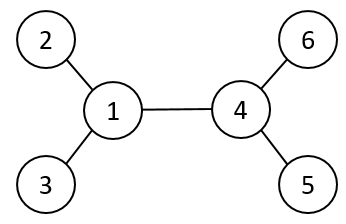
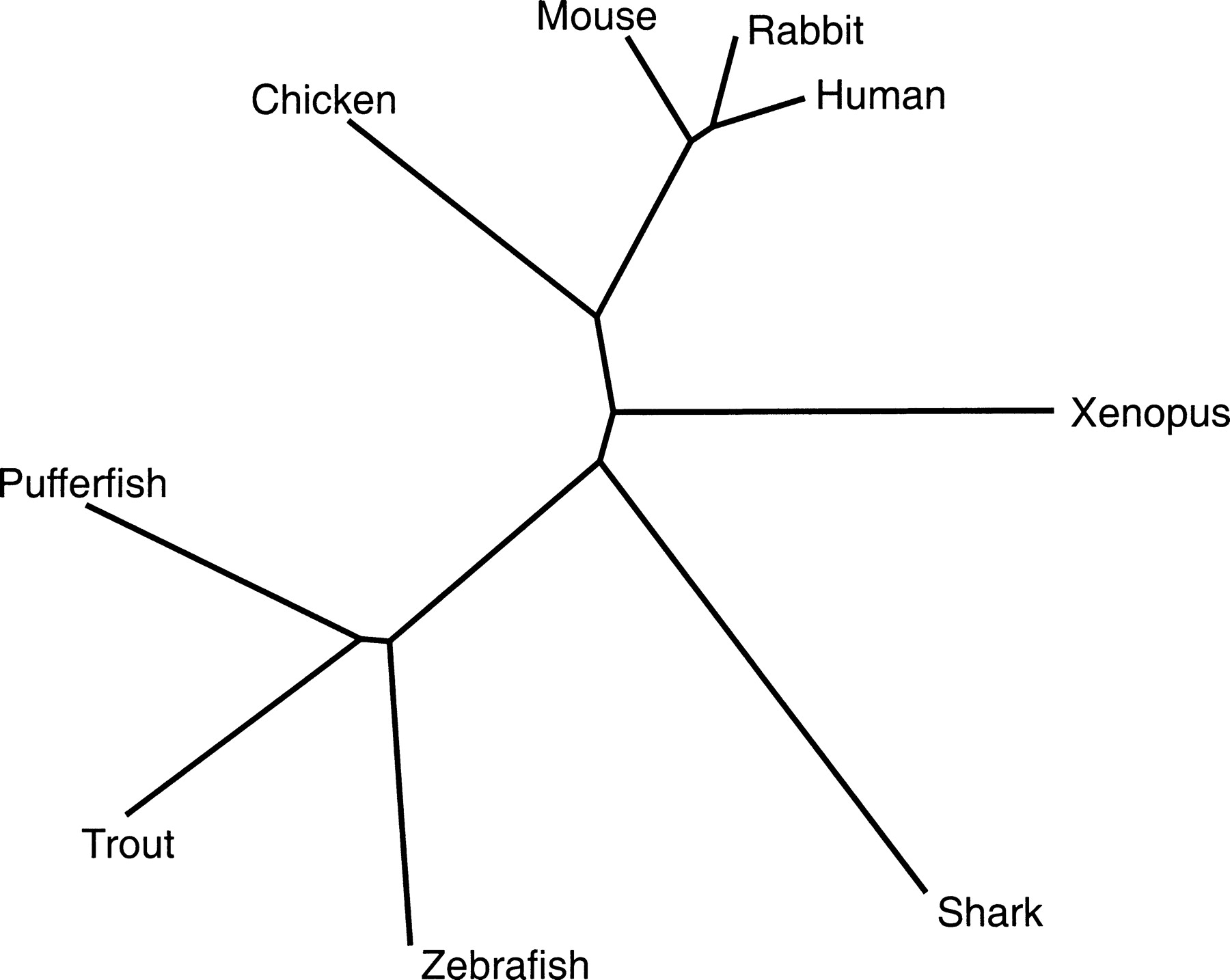
Unrooted Evolutionary Trees
When constructing evolutionary trees for diverse organisms, scientists often lack information about the “true ancestor” or the exact starting point of a lineage. In these cases, an unrooted tree is used.
- Relational Information: Although there is no “top-down” hierarchy, the tree visualizes the biological distance between species.
- Closeness vs. Difference:
- Leaves closer together on the tree branches are more biologically or genetically similar.
- Leaves farther apart represent organisms that are more biologically distinct.
- No Root: Because there is no assumed ancestor (root node), the tree focuses entirely on the relationships between the observed organisms rather than the chronological order of their evolution.
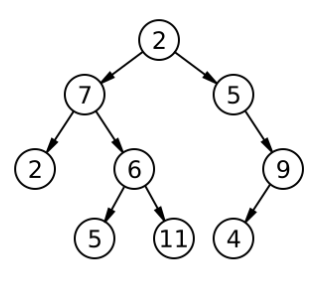
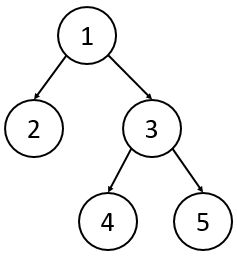
4. Rooted Binary Trees
A Binary Tree is a rooted tree with two specific restrictions:
- Every node except the root has exactly one parent.
- Every node has at most two children (commonly referred to as the “left” and “right” child).
5. Binary Tree Traversals
To access data in a tree, we typically start at the root and follow a traversal algorithm. There are four primary methods.
Consider a tree where V = Visit, L = Left Child, and R = Right Child:
| Traversal | Logic | Description |
|---|---|---|
| Pre-order | VLR | Visit node, then traverse Left, then Right. |
| In-order | LVR | Traverse Left, visit node, then traverse Right. |
| Post-order | LRV | Traverse Left, then Right, then visit node. |
| Level-order | BFS | Visit nodes level-by-level, left-to-right. |
All traversals’ worst case time complexity to traverse a binary tree of nodes =