ABSTRACT
In molecular biology, finding restriction enzyme motifs in a genome is a massive string-matching problem. While naive algorithms take time, the Aho-Corasick Algorithm uses a specialized finite state machine called an Automaton to find all occurrences of all motifs in a single linear scan.
1. The Scaling Problem
Searching for millions of short motifs () within a 3-billion-base genome () is computationally expensive.
- Naive Search: . Each motif is searched individually across every position in the genome.
// Naive Search
for each word w in D:
for each valid start position i of Q:
if w == Q's substring of length |w| starting at i:
w was found at position i of Q- Multiway Trie: . By combining motifs into a tree, we check multiple words simultaneously, but we still have to restart the search for every new starting position in the genome.
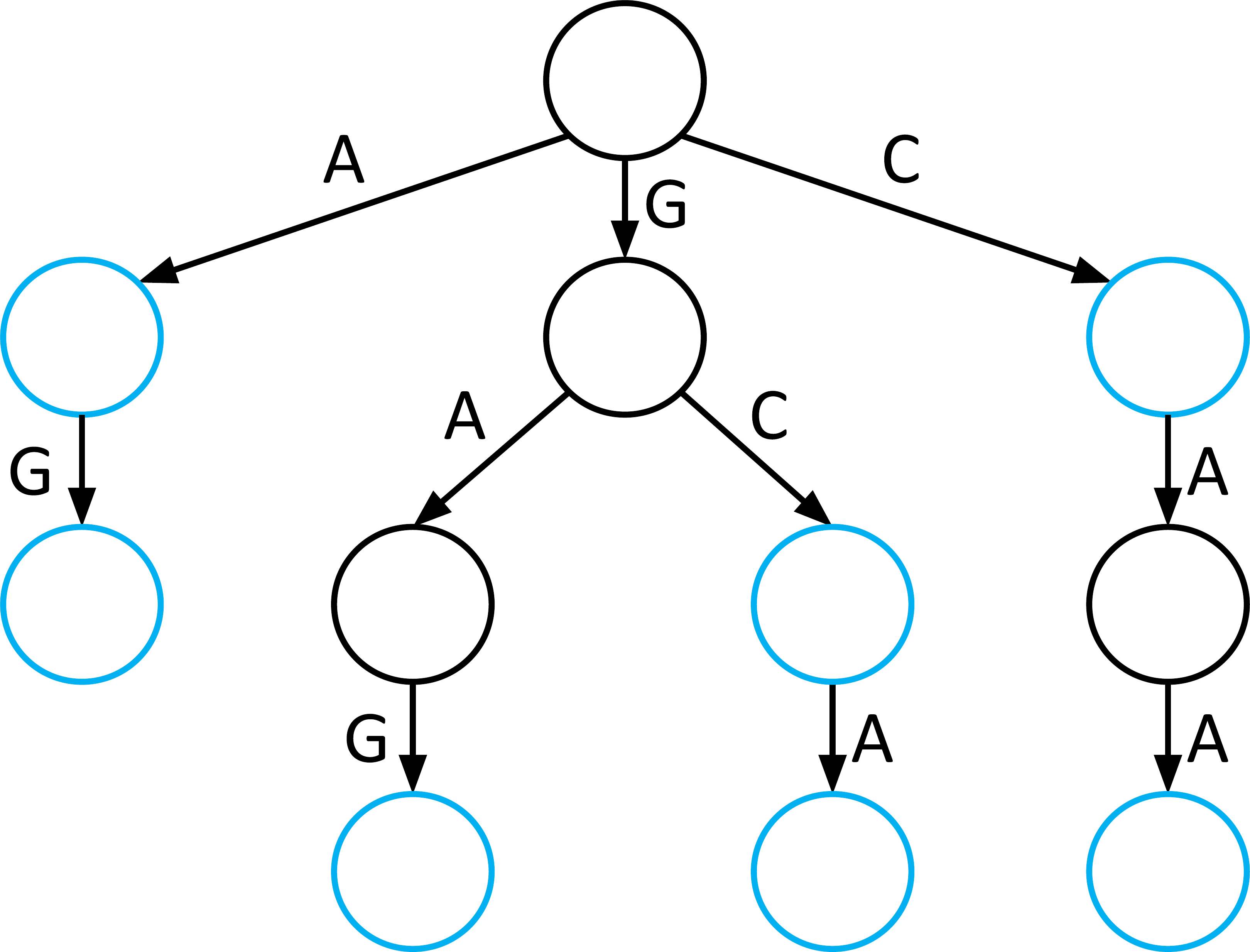
2. The Aho-Corasick Automaton
The Aho-Corasick Automaton solves the restart problem by adding “shortcuts” to a standard Trie. This allows the algorithm to transition between motifs without ever re-reading a character of the genome.
Failure Links: The Error Recovery
A Failure Link connects a node to another node if the path to is the longest possible suffix of the path to .
- If you fail: When the next character in the genome doesn’t match any child edge, you follow the failure link.
- Why it works: It preserves the progress you’ve already made by jumping to the start of another word that shares a suffix with what you just typed.
Not including the full path from the root to
This will cause the failure link to always point to
for each node 'curr' in a BFS starting at the root:
if curr is the root or is a child of the root:
create failure link from curr to the root
else:
p = parent of curr
c = label of edge going into curr
x = node pointed to by p's failure link
repeat infinitely:
if x has a child with edge label c:
create failure link from curr to that child of x
break
else if x is the root:
create failure link from curr to the root
break
else:
x = node pointed to by x's failure link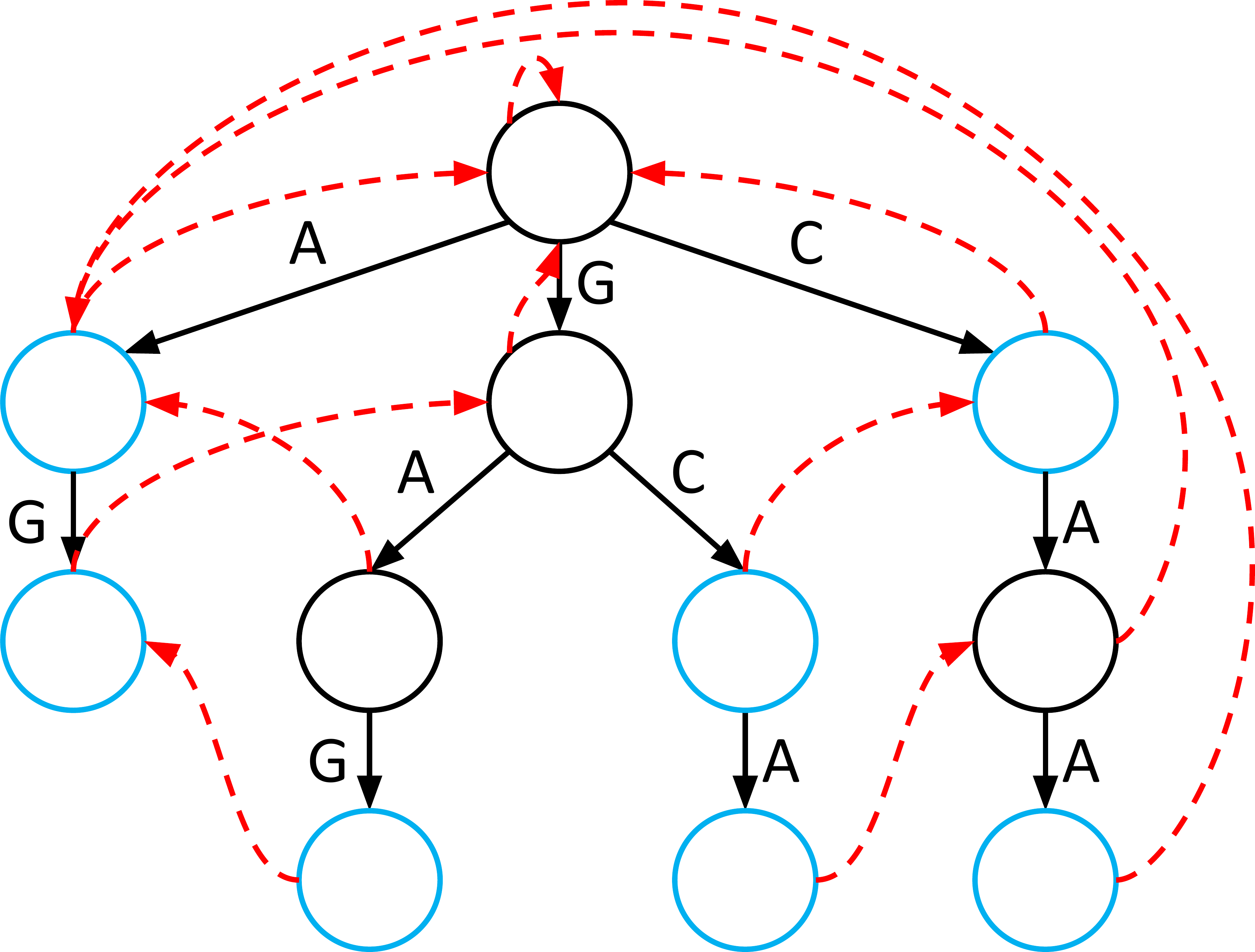
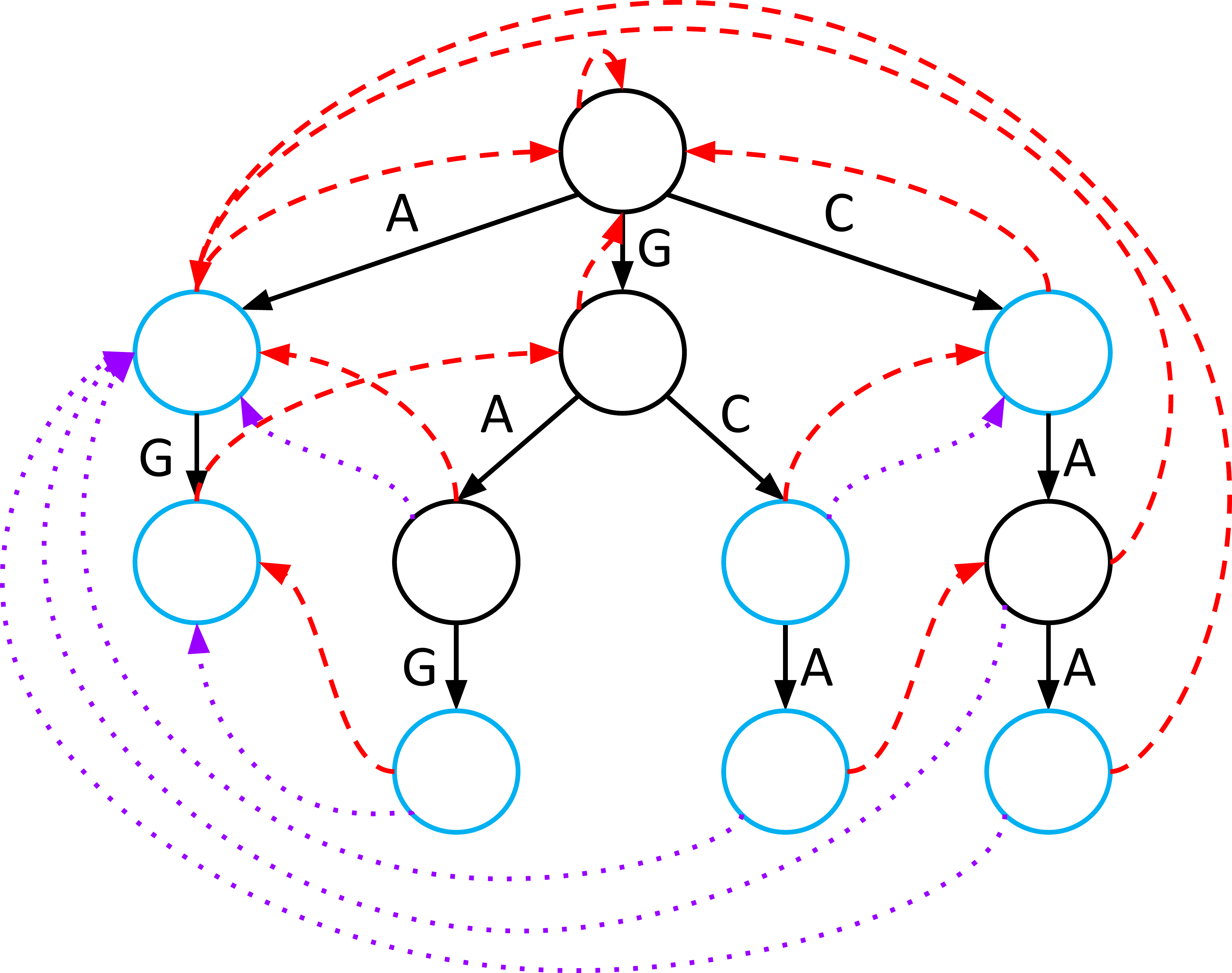
Dictionary Links: Finding Hidden Words
Sometimes, a word exists completely inside another word (e.g., “A” exists inside “GCA”). If you are at the end of “GCA,” you might miss “A” because it ended earlier.
- Dictionary Links point to the nearest node that represents a complete word in your database.
- When you land on a node, you follow its dictionary links to report every word that ends at that current position in the genome.
- For each node , draw a link to the first word node you would encounter if you were to repeatedly traverse failure links. If no such word node exists, will not have a dictionary link.
3. Algorithm Summary
Construction (Preprocessing)
- Build a Multiway Trie of all motifs in your database.
- Use a Breadth-First Search (BFS) to calculate failure links.
- Calculate dictionary links to ensure no overlapping motifs are missed.
Scanning (The Linear Scan)
The scan is because the pointer in the genome only ever moves forward. If a mismatch occurs, the “state” of the automaton shifts via failure links, but the genome index remains the same.
scan(genome):
curr = root
for each nucleotide c in genome:
while curr cannot move to c:
if curr == root: break
curr = follow_failure_link(curr)
curr = move_to_child(curr, c)
// Report every motif found ending at this position
report_all_matches(curr) 4. Performance Comparison
| Algorithm | Complexity | Efficiency |
|---|---|---|
| Naive Search | Extremely Slow | |
| Multiway Trie | Much Faster | |
| Aho-Corasick | Optimal |